Research Themes
Social behaviors are diverse in nature, but it remains unclear how conserved genes, brain regions, and cell populations generate this diversity. I study how this diversity arises from evolutionarily conserved gene regulatory features and cellular states of the vertebrate brain, examining how gene expression, chromatin accessibility, and spatial organization of cell types collectively shape neural circuits and behavior through single-cell neurogenomics. I develop statistical modeling and machine learning approaches to turn rich, high-throughput, high-content biological measurements into quantitative, testable theories and useful computational tools.
Brain, Behavior, and Evolution
Context-dependent processes like embryonic development, the regeneration of organs and complex behavior are fascinating because they reveal new rules of biological systems that are not necessarily operational during homeostasis. For instance, recent results suggest that stem-like cells in the brain may tune the evolution of male social behavior. Using single-cell multi-omic profiling of the vertebrate forebrain, I investigate how brain circuits are rewired, how the composition of specific cell populations is rebalanced, and how epigenomic information and cis-regulatory enhancer–gene mapping are dynamically reconfigured across behavioral states.
Single-Cell & Spatial Omics
Single-cell and spatial omics provide unprecedented resolution to reconstruct cell states, lineage relationships, and tissue architecture in situ. I develop and apply multi-omic analytic frameworks to map cell-type diversity, resolve how behavioral state reshapes cellular programs over time, and link cis-regulatory elements to gene expression within spatial context. Through these efforts, I aim to uncover how spatial organization and regulatory landscapes coordinate cellular function across development, regeneration, and behavior.
Statistical Learning & Modeling
While I rarely build new tools for their own sake, single-cell data from non-model species is heterogeneous, high-dimensional, and structured in ways that off-the-shelf methods struggle to handle. To extract interpretable understanding from the data, I develop approaches that combine graph-based statistical modeling with modern machine learning, spanning latent representation learning, deep learning characterization of enhancer code evolution, and probabilistic methods for cell-type trajectory inference, causal GRN modeling, cell phylogeny reconstruction, and allelic imbalance detection.
Selected Publications
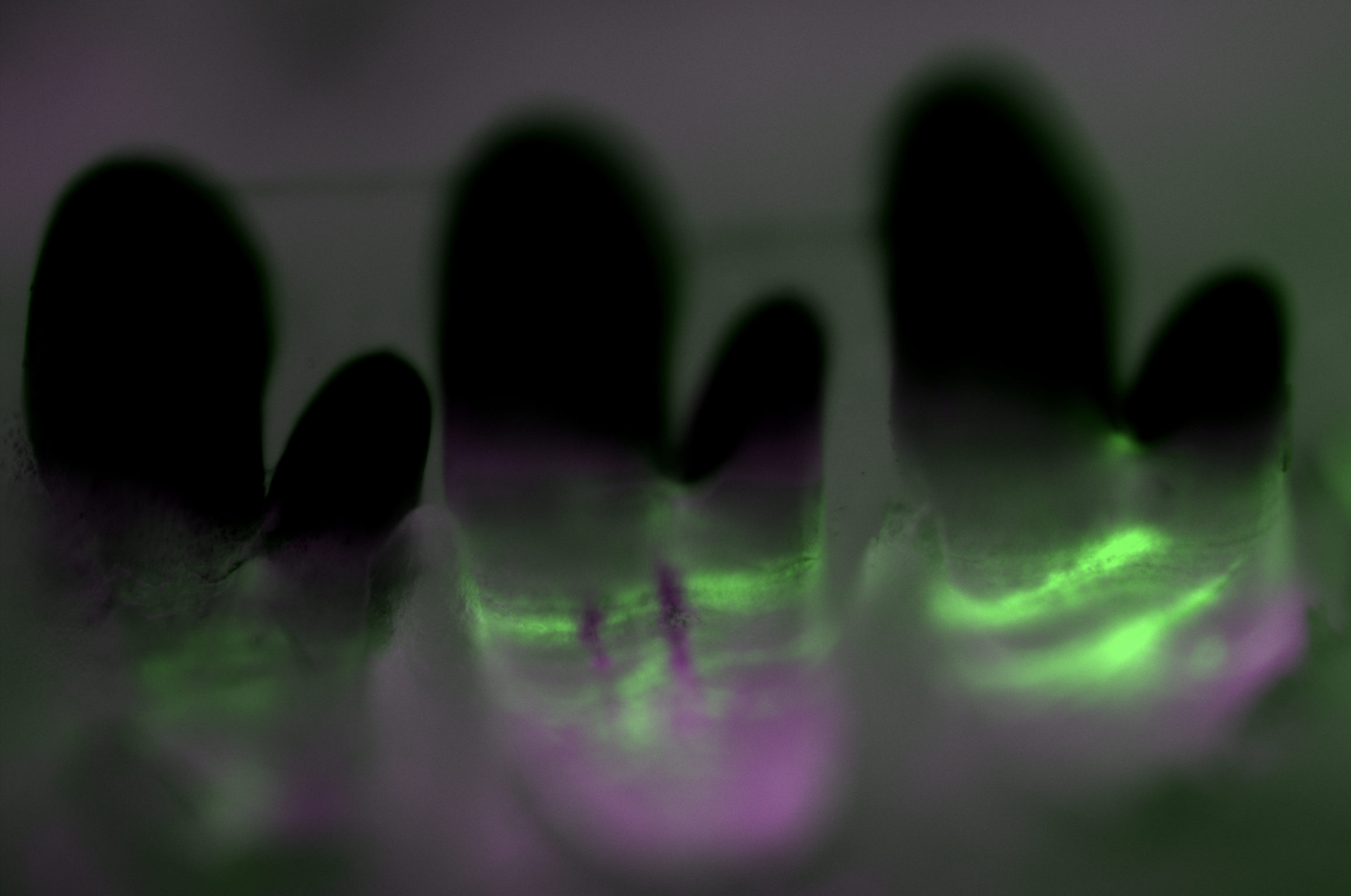
Cellular basis of accelerated whole-tooth regeneration
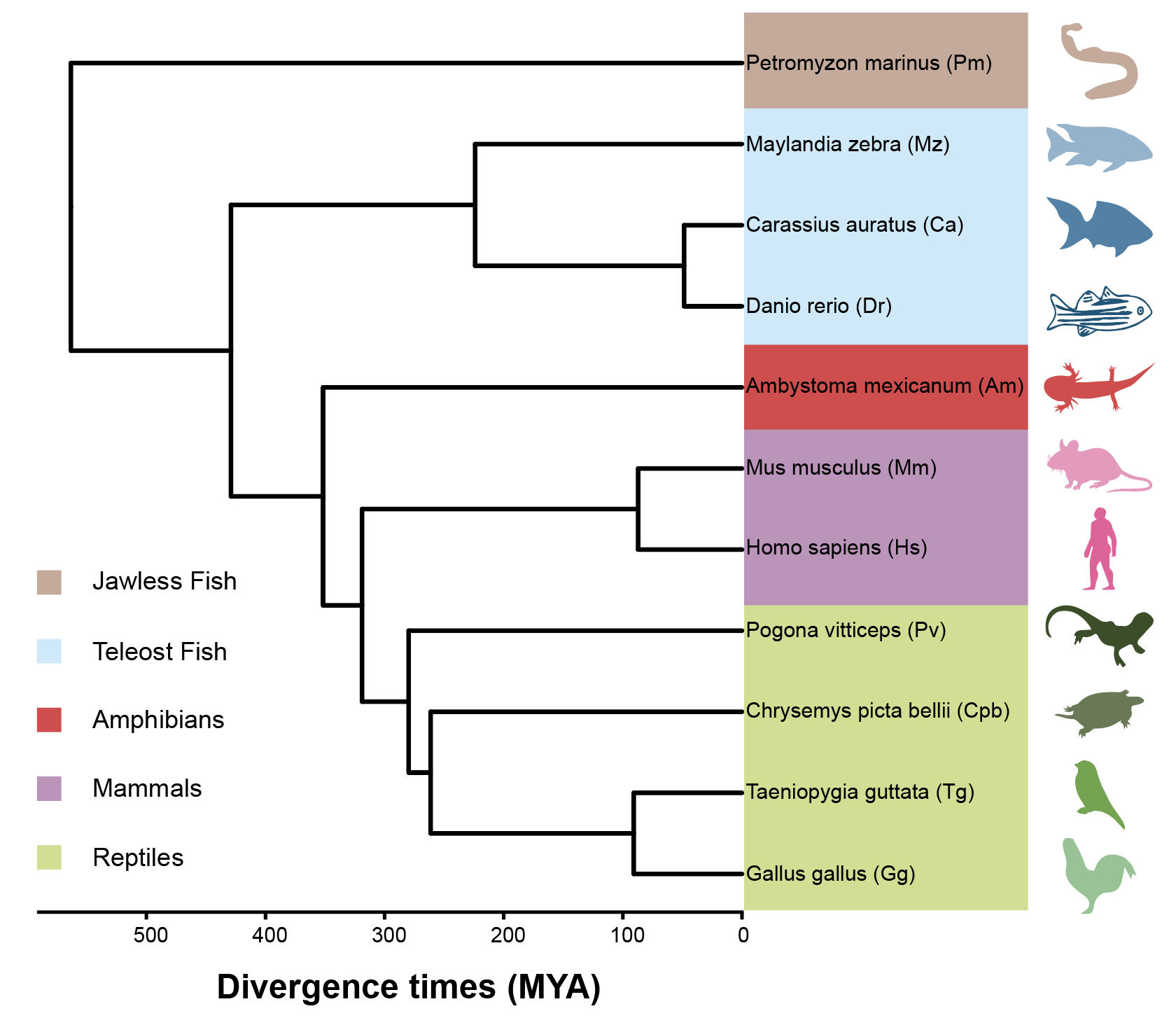
Evolutionarily informed gene sets reveal conserved and lineage-modified transcriptional programs during vertebrate forebrain evolution
The Team

Jacy Zanussi
stochastic gene expression + simulation-based inference
fav veggie: all peppers 🌶️ (approved as honorary vegetable)

Austin Marcus
nuclear condensates, cilia (co-advised w/ Jun Allard)
fav veggie: onion 🧅

Rebecca (Rujie) Gu
physics-informed neural networks + point processes
fav veggie: spinach 🍃

Matt Lastner
meiosis biophysics, polymer models (co-advised w/ Ofer Rog)
fav veggie: sweet potato 🍠
You?
Prospective students can apply to the Utah math graduate program and/or reach out to Chris. Be prepared to defend your favorite vegetable.
Apply to Utah MathAbout Chris
Timeline





Editorial


Miscellany

Chris + little mathematicians
As a first-gen college student, I care about outreach and inclusion in science, especially for historically marginalized populations. Some programs I've been part of: Science Communication Fellow (Utah), INSPIRE (incarcerated youth), Proud to be First (NYU), Cal-Bridge, and DECADE (UCI).

Finn (left) + Piper (right) + foster (middle)
In rare instances of spare time, you might find me: birding, rock climbing, watching arthouse film, or tending to the ever-growing needs of my cats (see photo). Some recent favorites include the American Kestrel, Wong Kar-wai's In the Mood for Love, and the humble beet .
Chris Miles (artist) + Chris Miles (mathematician)
For the sake of clarity: I am not Chris Miles, the rapper. Nor am I Christopher J. Miles, the other scientist. I am also not Chris Miles, the artist. However, I have crossed paths with him and appreciate his work.














